Biology, also known as life science, is one of the core
branches of knowledge. It deals with the vital processes of living
organisms. The history of research and development in this field is
quite ancient. With the development of computer technology, men
have created some real progress in this field. From conquering
fatal diseases to solving the mystery of a living organism, the
computer is a great companion for the biologists. There are many
open-source biology tools available out there. Linux is a very
customizable open-source operating system that is preferred by many
researchers. So if you are a biologist or an amateur biology
enthusiast looking for some Linux biology software, you might want
to check out these biology tools for Linux PC to get the most out
of your study or research.
Some people have a common misconception that Linux doesn’t
have a huge library of software. But you will be surprised that in
the category of education and research-based software, Linux is
still unbeatable. It’s because most of the scientists and
researchers are with the open-source software movement.
Hence you are getting an extensive collection of biology
tools for Linux. They are free and no less than any paid software.
Here I have made a curated list of 15 tools of different types so
that you don’t need to take any hassle finding them. If you go
through this full article, I hope you will find the best software
for the Linux system, which will suit your needs.
1. EMBOSS
The explication of the name of the software is
the European Molecular Biology Open Software Suite. It is an
open-source biology tool for Linux made for interested people in
the field of biology. EMBOSS is a powerfull sequential analysis
tool. It is somewhat a complete package of tools that the features
and possibilities of this tool are beyond explanation.
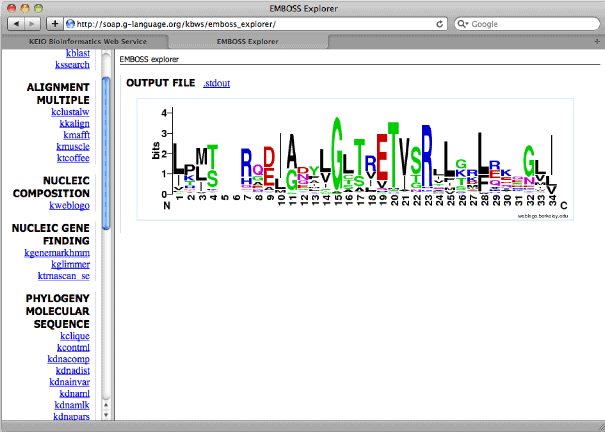
Key Features of EMBOSS
- It can rapidly crawl and retrieve sequential data from
the web. - EMBOSS is used for sequence alignment, protein motif
identification, nucleotide sequence pattern analysis,
etc. - It has a built-in library for releasing new open-source
tools. - An advanced presentation tool is built-in with this for
quick publication of the retrieved data. - It can do string handling, pattern-matching, list
processing, and database indexing by using additional programming
libraries. - The integration feature is useful for syncing with other
popular tools.
2. NAMD
NAMD is a simulation program developed especially for simulating
huge biomolecular systems. This biology tool for Linux is so
powerful that it can process millions of atoms at a time
parallelly. Charm++ is a C++ based language that is used to write
this program. NAMD uses a runtime environment named Converse to run
on parallel cluster-based systems, which helps to process huge
amounts of biological data at a time.
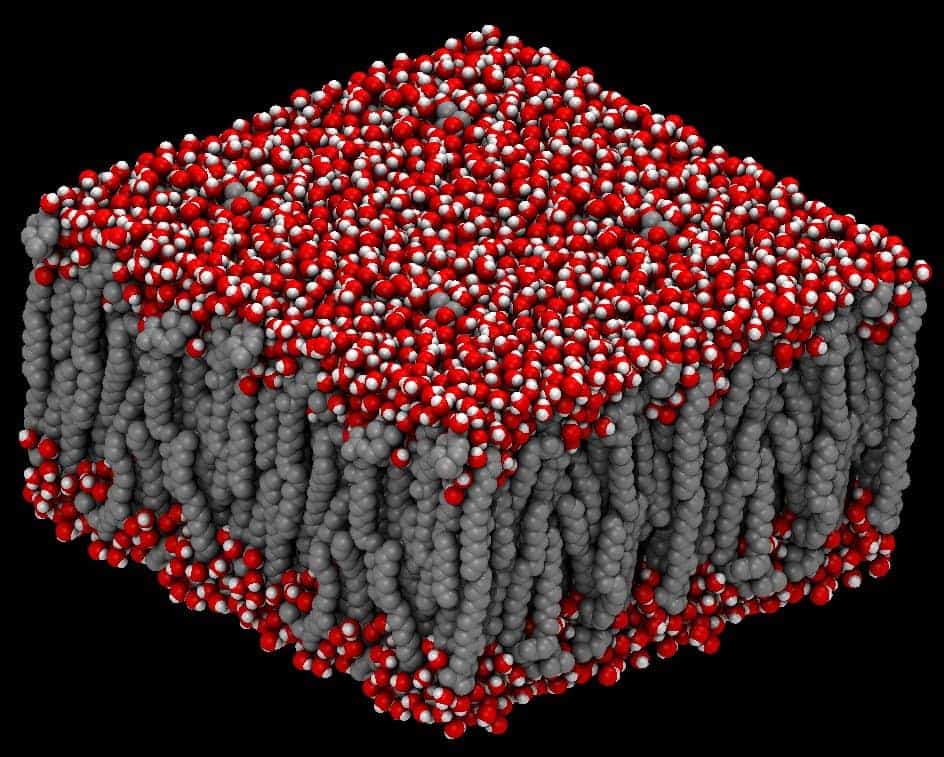

Key Features of NAMD
- Molecular structure simulation is prepared by using Visual
Molecular Dynamics. - It supports different types of input files, including X-PLOR,
CHARMM, AMBER, etc. - NAMD uses multi-time-step integration for numerical
analysis. - Users can select from a wide range of dynamics simulation
options. - It supports GPU accelerated processing.
- This tool supports Replica-based umbrella sampling via the
collective variables module.
3. GROMACS
GROMACS is not only yet another biological simulation tool;
rather, it is a complete software package with integrated building
and analysis tools. This versatile biology tool for Linux can
perform analysis and simulation for thousands to millions of
biological particles. It was primarily developed for the analysis
of biological chemicals like proteins and lipids. But now it is
also used in non-biological research fields.

Key Features of GROMACS
- This tool is two to three times faster than it’s
competitors. - The software code is highly optimized for faster data
processing. - Gromacs is pretty user-friendly. The error codes are written
with plain texts for easier understanding. - The extensive user manual for this tool is available for free
of cost in e-paper format. - It can store trajectory data in a compact method.
- It has some integrated tools for trajectory analysis. Users
don’t need to write any codes for this purpose. - It features a fully automated topology builder for proteins,
which is very useful.
4. VMD
VMD is an advanced bio-molecular visualization program developed
for Linux. A molecular visualization program is mainly a program
for displaying molecular data with 3D graphics. VMD can read and
analyze PDB or Protein Data Bank files and render them in a
structured graphical manner. It can even simulate molecules for
different conditions and cases. Thus it has become a very useful
program for the deep researchers of biology.
Key Features of VMD
- It can utilize the external GPU power of the computer.
- The developer has not applied any limitations for the number of
molecules or other parameters. The RAM is your limit! - Users can easily generate PDF files from the standard 3D output
with the built-in tool. - VMD can utilize the stereo display system provided you have
that. - The extensive library of built-in file reader supports up to 60
different file formats. - Researchers can write their routine commands using Tcl
language.
5. simuPOP
SimuPOP is not yet another ordinary biology tool for Linux.
Rather it is a forward-time population genetics simulation
environment. It can analyze and simulate any population-related
problems. Hence the researchers in the field of biology use this
tool for simulating the spreading of complex diseases. simuPOP uses
Python as a core scripting language.

Key Features of simuPOP
- It has the option to attach information fields to individuals
of a population. - It has number limits for the number of homologous sets of
chromosomes or other parameters. - It has more than 70 built-in operators for population
analysis. - The advanced scripting interface gives users the ability to
customize this program. - simuPOP has a comprehensive documentation system for
beginners.
6. MUSCLE
MUSCLE is the abbreviation for the original software name MUSCLE
MUltiple Sequence Comparison
by Log- Expectation. It
is a very popular biology tool for Linux, which is used for
creating multiple alignments of amino acid or nucleotide sequences.
Besides, it’s better accuracy, and better speed keeps it ahead of
the other competitors like ClustalW2 or T-Coffee. It is considered
as one of the fastest programs in this category.
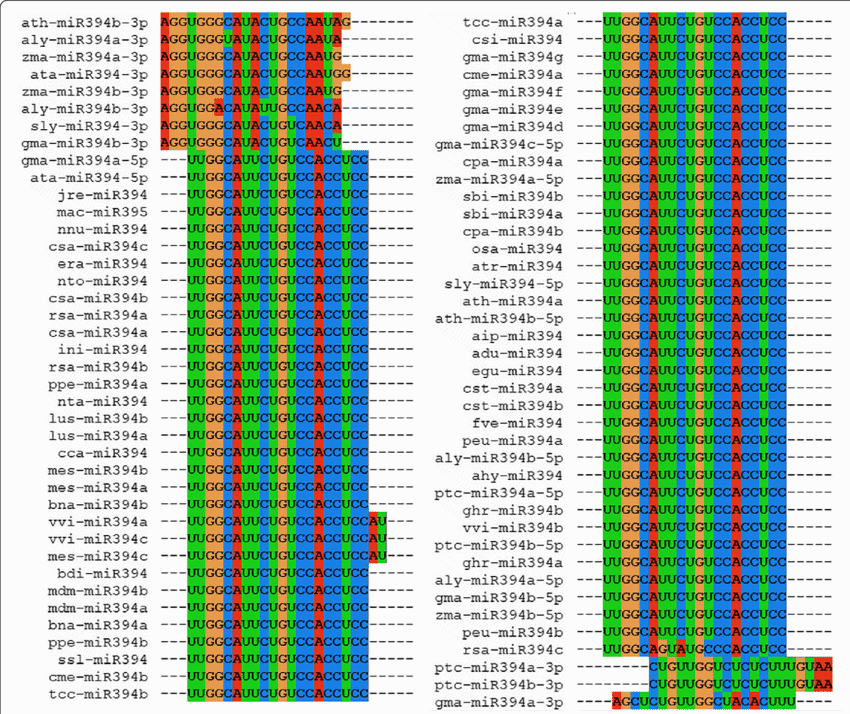

Key Features of MUSCLE
- It supports three different protein profile scoring
functions. - MUSCLE provides diagonal and anchoring optimization
features. - The popular text-based format FASTA is used in this tool as
both input and output files. - It features an extra benefit that can generate output files as
different popular formats like LUSTALW, MSF, HTML, etc.
7. SeaView
SeaView is a normal multiple sequence alignment software. But
it’s specialty is that it has a very good and easy to use graphical
user interface. This package is used as the backend for different
other popular tools like Clustal Omega, Gblocks, and PhyML. Fast
Light Toolkit, commonly known as FLTK powers the user interface of
this program.

Key Features of SeaView
- It supports most file formats for DNA and protein sequencing,
including NEXUS, MSF, CLUSTAL, FASTA, PHYLIP, etc. - Users can import external FASTA format files for alignment
algorithms. - It can draw phylogenic trees and generate them in different
common formats like PDF, SVG, EPS, etc. for printing or
publishing. - SeaView has embedded downloader for downloading genetic
sequence from the internet.
8. TREE-PUZZLE
TREE-PUZZLE is the new name for the software PUZZLE. It is a
very popular biology tool for Linux. But originally, it is a
console-based tree search algorithm that is used for the analysis
of large data sets. This TREE-PUZZLE software package can
reconstruct trees by using the algorithms described by Strimmer and
von Haeseler.
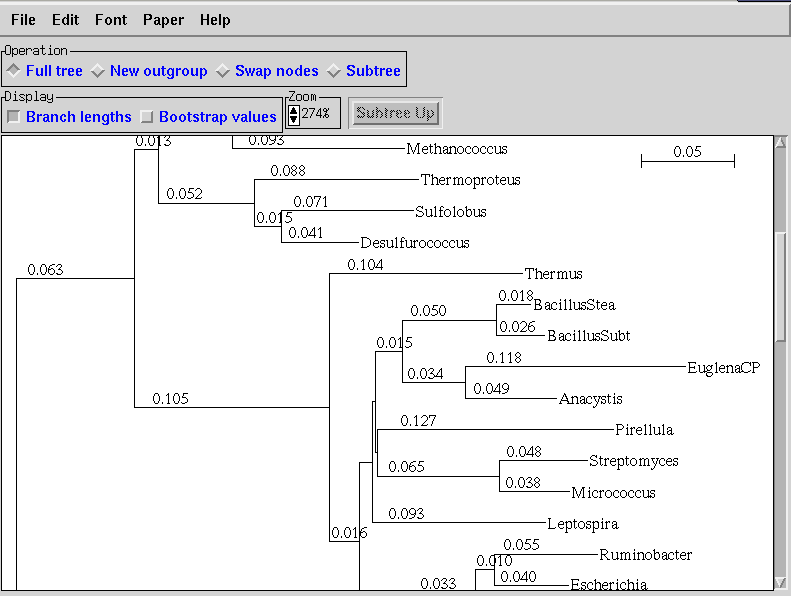
Key Features of TREE-PUZZLE
- It uses quartet puzzling algorithms.
- This tool can assign estimations of support for each internal
branch automatically. - TREE-PUZZLE can construct trees by inputting user given sets of
trees. - It has some tools to conduct statistical tests on the data
sets. - It can estimate parameters and pairwise distances.
9. TreeView X
It is an open-source biology tool for constructing phylogenic
trees. Tree construction software is very important in the field of
biology. That is why it is considered as a good Linux biology tool.
It can read tree files with different file formats.
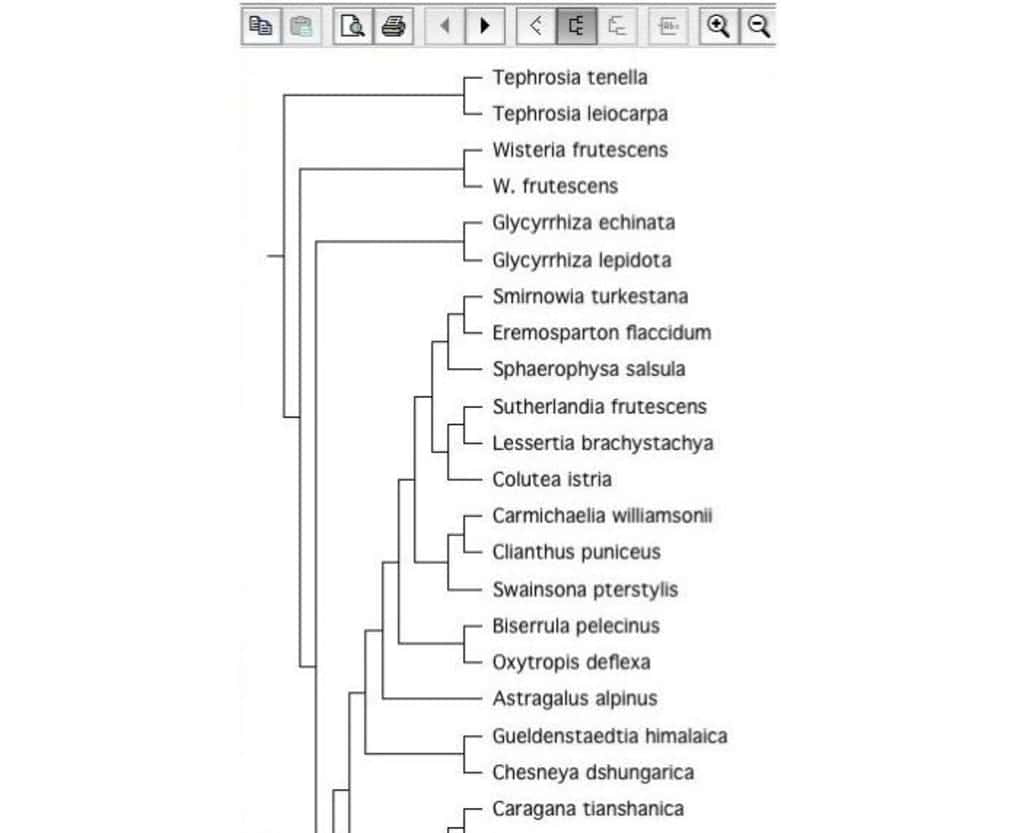
Key Features of TreeView X
- It has a rich GUI based on the wxWidgets C++ library.
- It can export trees in different image-based file formats.
- TreeView X has an advanced printing option built-in that helps
in formating printing paper numbers according to the user’s
need. - The drag and drop feature increases productivity while using
this tool.
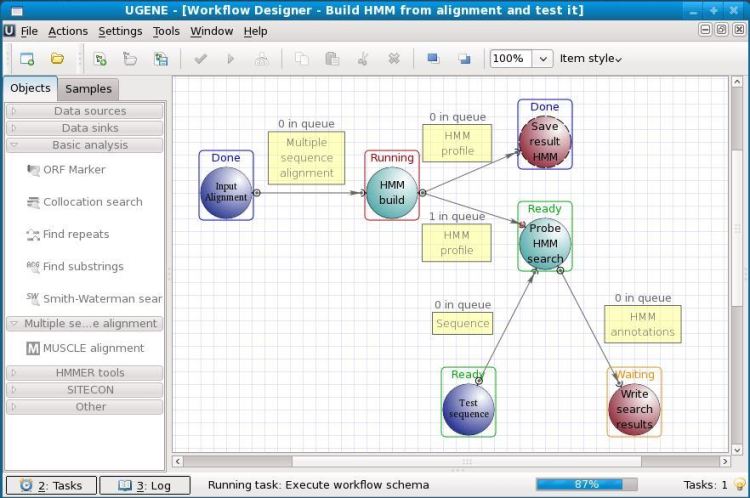
10. UGENE
It is an open-source biology software for Linux. UGENE is used
for the analysis of various biological data. Nowadays, it is mostly
used for genome sequencing. The analyzed data can be stored on the
computer storage or even on a shared lab database. The graphical
user interface of this tool helps users to operate this without any
prior coding knowledge. Other than GUI, it also has a legacy
command-line interface to work with.
Key Features of UGENE
- Users can create and annotate protein sequences easily.
- It can utilize the multiple cores of the host CPU and can
utilize a discrete graphics card. - It has built-in integration with popular bioinformatics servers
like PDB, NCBI, etc. - UGENE has an integrated Primer3 tool for the design of a PCR
primer. - It features an advanced chromatogram viewer.
- This tool can search for complex signals with
ExpertDiscovery.
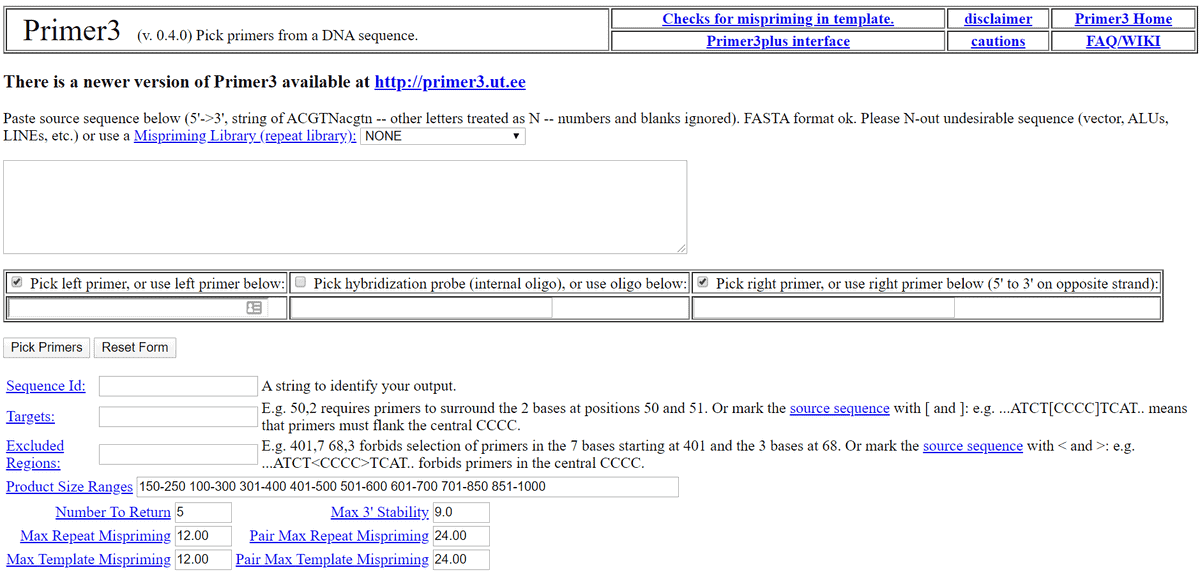
11. Primer3
Primer3 is one of the most popular biology software for Linux.
It is a free and open-source biology tool for Linux under the GNU
license. This tool is used for picking the primer from a DNA
sequence. This tool also has an alternative web user interface
named Primer3 Plus for those who don’t want to install it
locally.
Key Features of Primer3
- Users van import/upload sequence files in almost any popular
file format. - Sequences can be pasted in plain text.
- It has many customization features under the general and
advanced settings category. - Users can input sequence quality in this tool.
- There is a dedicated tab for penalty weights in this tool.
12. Integrated Genome
Browser
As the name suggests, it is a genome browser for your desktop.
It is a free and open-source biology tool. This biology software
for Linux can search for genome sequences from the internet. Of
course, you can search for these particular bioinformatics data
through your regular browser. But trust me, this dedicated browser
will make your workflow a lot faster. This tool is built upon the
Genoviz SDK, a Java library.

Key Features of Integrated Genome
Browser
- This tool can read data from many file-formats, including BAM,
BED, Affymetrix CHP, FASTA, GTF, PSL, etc. - Users can export the output to any printable formats like SVG,
PNG, or even easy to use PDF. - Dynamic and real-time zooming and scrolling features.
- It supports REST-style web services for annotation
features.
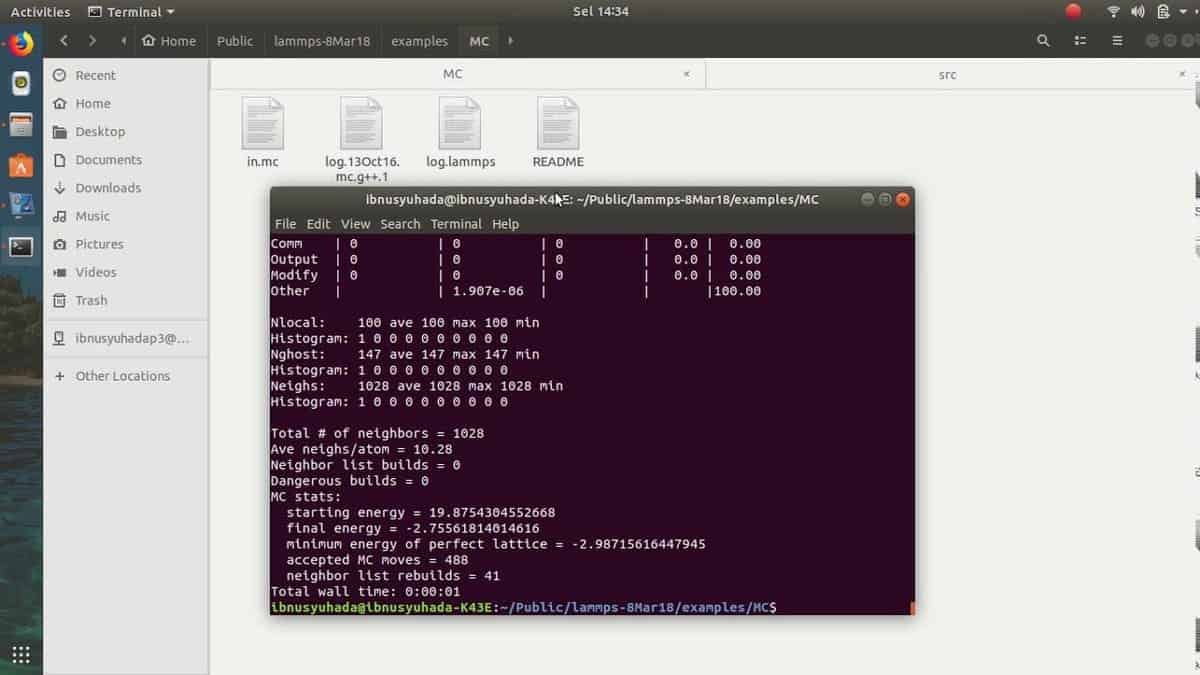
13. LAMMPS
LAMMPS is one of the most popular open-source biology tools. The
abbreviation stands for “Large-scale
Atomic/Molecular
Massively Parallel
Simulator.” It is a general-purpose molecular
dynamics software. But nowadays, it is highly used in the field of
biological research. It is developed and maintained by Sandia
National Laboratories. This Linux biology software uses Message
Passing Interface or MPI protocol for parallel communication among
researchers.
Key Features of LAMMPS
- It uses an efficient data structure named the Verlet List for
keeping track of nearby particles. - It can utilize the full potential of a parallel computing
system by dividing the simulation domain into smaller subdomains
and distributing them for each processor. - This tool is highly portable because it is made upon C++.
- Built-in support for CUDA and OpenCL GPU rendering system.
- Users can easily extend new features and functions.
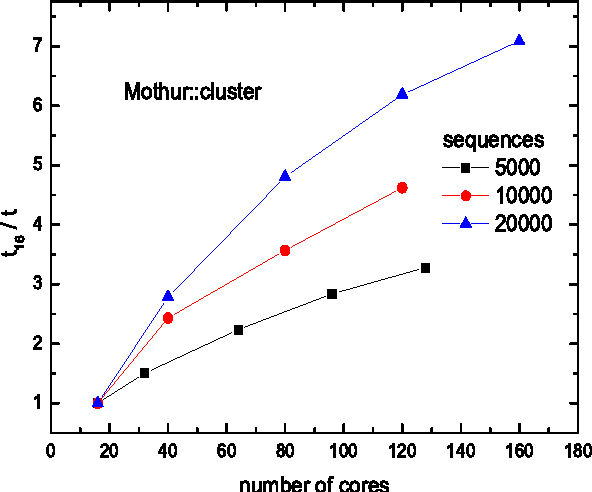
14. Mothur
Mothur is a well-known Linux biology software among scholars.
This software project was initiated by Dr. Patrick Schloss et al.
Many publications of biological research cited this software so
far. This open-source tool is a very efficient bioinformatics data processor[14]. It is mostly used for
the DNA analysis of the uncultured microbes.
Key Features of Mothur
- It can process data generated from several DNA sequencing
methods. - Almost all the popular methods are supported by this tool,
including 454 pyrosequencing, Illumina HiSeq and MiSeq, Sanger,
PacBio, and IonTorrent. - No other tools can beat Mothur in analyzing 16S rRNA gene
sequences. - It is maintained regularly by a group of well-known scholars of
biology.
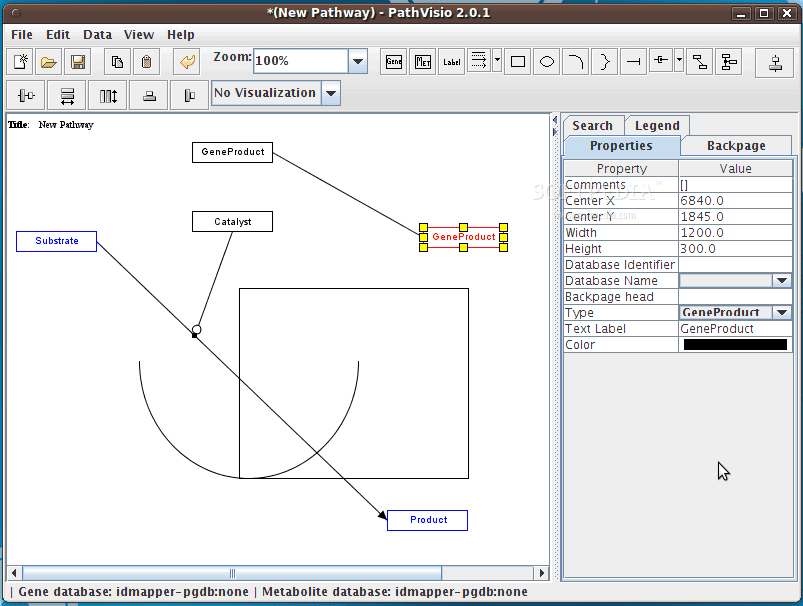
15. PathVisio
PathVisio is a free and open-source biology tool for Linux. It
is used for drawing, editing, and analyzing biological pathways. It
has many useful features built-in with the package. Users can also
install additional features via plugins. This tool is based upon
Java, and this is why it can easily be installed on any platform,
including Linux.
Key Features of PathVisio
- Advanced drawing and annotation tools for pathways.
- It can even analyze different types of biological
pathways. - PathVisio has built-in integration with WikiPathways for easier
publishing. - The open-source tool Cytoscape can be easily integrated with
this tool. - It can be integrated with other programming languages via
PathVisioRPC.
Final Thoughts
As you can see, there are numerous tools for the different
purposes needed in the field of biology. Biology is a vast field of
knowledge and research. So it is obvious that you won’t need to use
all of the tools mentioned above. If you try out this curated list
of Linux biology software, you will get to know which will suit
your works best. And, if you have any favorite software in this
category, you can let others know by commenting below.
References
- ^
Get EMBOSS
(emboss.sourceforge.net) - ^
Get NAMD
(www.ks.uiuc.edu) - ^
Get GROMACS
(www.gromacs.org) - ^
Get VMD
(www.ks.uiuc.edu) - ^
Get simuPOP
(simupop.sourceforge.net) - ^
Get MUSCLE
(www.drive5.com) - ^
Get SeaView
(doua.prabi.fr) - ^
Get TREE-PUZZLE
(www.tree-puzzle.de) - ^
Get
TreeView X (code.google.com) - ^
Get UGENE
(ugene.net) - ^
Get Primer3
(primer3.ut.ee) - ^
Get IGB
(bioviz.org) - ^
Get
LAMMPS (lammps.sandia.gov) - ^
Top 20
Best Bioinformatics Tools for Linux: An Ultimate Collection
(www.ubuntupit.com) - ^
Get Mothur
(www.mothur.org) - ^
Get PathVisio
(pathvisio.github.io)
Read more https://www.ubuntupit.com/best-biology-tools-for-linux/